For the dietary supplement market, we have created a range of unique high-quality natural ingredients whose innovative nature has been scientifically substantiated. All of Our ingredients are certified to applicable organic standards: National Organic Program (NOP) and European Union (EU), in order to ensure that your product benefits from optimal positioning, to keep you ahead of the competition in the dietary supplement market. Astragalus Extract,Organic Astragalus Root Slices,Astragalus Root,Organic Astragalus Root Organicway (xi'an) Food Ingredients Inc. , https://www.organic-powders.com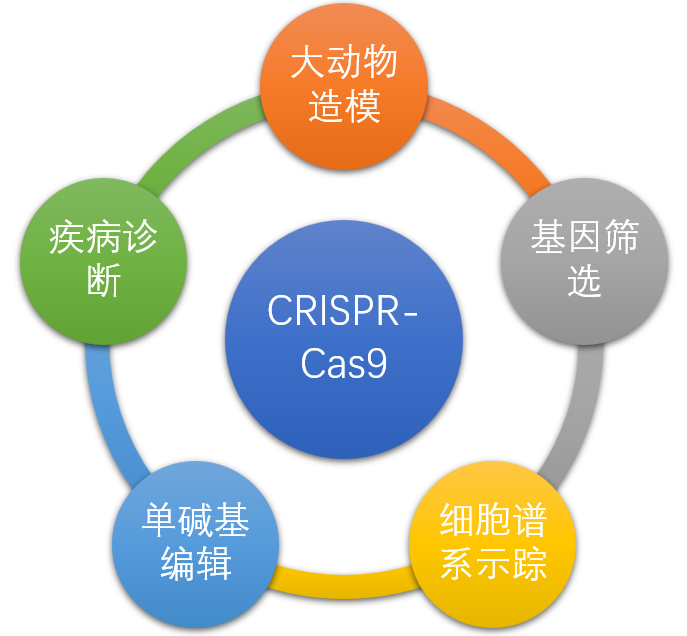
Five application areas of CRISPR technology
First, large animal modeling
The CRISPR-Cas9 system provides an efficient method for the construction of animal models with its simple operation. At present, there are thousands of animal models of human diseases based on CRISPR-Cas9, mainly in the rodent (such as mouse) model, and its related pathogenesis and treatment progress provide many valuable clinical applications. reference. Bangyao Lab has also been committed to related research, published many articles on the construction of gene-editing large mice and its application in various diseases in internationally renowned journals, and provided hundreds of genetically edited animals for the majority of scientific users. model.
Although the mouse model can well demonstrate the characterization of the disease, due to the huge differences in lifespan and physiological structure, many neurodegenerative diseases and age-related diseases do not mimic the corresponding effects of many human diseases. Characterization. Therefore, the construction of large mammalian models that are closer to humans in genetics, anatomy, and physiology is particularly important. Due to the difficulties of embryonic stem cell technology, long reproductive cycle and low efficiency of gene editing, the construction of large mammalian models has always been a difficult point in scientific research.
However, with the rapid development of CRISPR-Cas9 technology, existing gene editing tools have been proven to successfully construct large mammalian models. In 2018, Chinese scientists made important contributions to genetically edited pigs and monkeys , leading the world in the construction of genetically edited large mammalian models based on CRISPR-Cas9 technology.
1. "Knock-in" pig: the construction of Huntington's disease pig model
In March 2018, several teams from China collaborated to establish a “knock-in†Huntington's disease (HD) porcine model, which uses CRISPR-Cas9 technology to integrate HD mHTT gene fragments into porcine fibroblasts. The mHTT transgenic pig model was obtained by somatic cell nuclear transfer technology, which provides an important animal model for testing HD therapy [2].
2. Six articles laid the world's leading position for scientists in China to build genetically edited monkey models
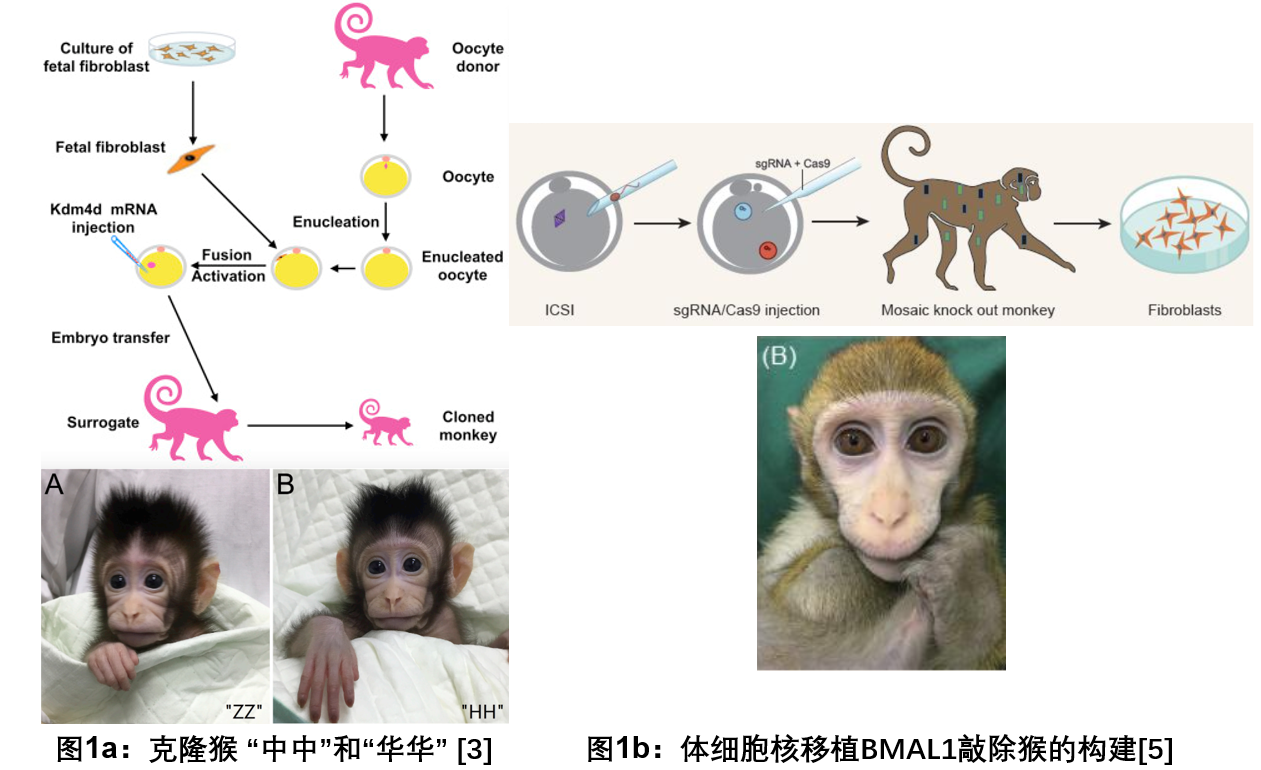
In early 2018, researchers at the Institute of Neuroscience of the Chinese Academy of Sciences reported for the first time that the limitations of somatic cell cloning of non-human primates were successfully overcome by removing chemical modifications on DNA when embryonic cells were differentiated into specific cells. The birth of a genetically cloned monkey was first realized (see Figure 1a), marking a milestone in non-primate somatic cell cloning technology [3].
l A year later, the research team published two articles in succession. The BMAL1 knockout monkeys with sleep disorders and mentally related abnormalities were constructed by CRISPR-Cas9 technology, and then the skin fibroblasts of BMAL1 knockout monkeys were used as nuclei. The donor has obtained a genetically consistent disease knockout monkey model by means of somatic cell nuclear transfer (see Figure 1b) [5], which lays a solid foundation for the construction of non-primate disease models and its application in clinical research. .
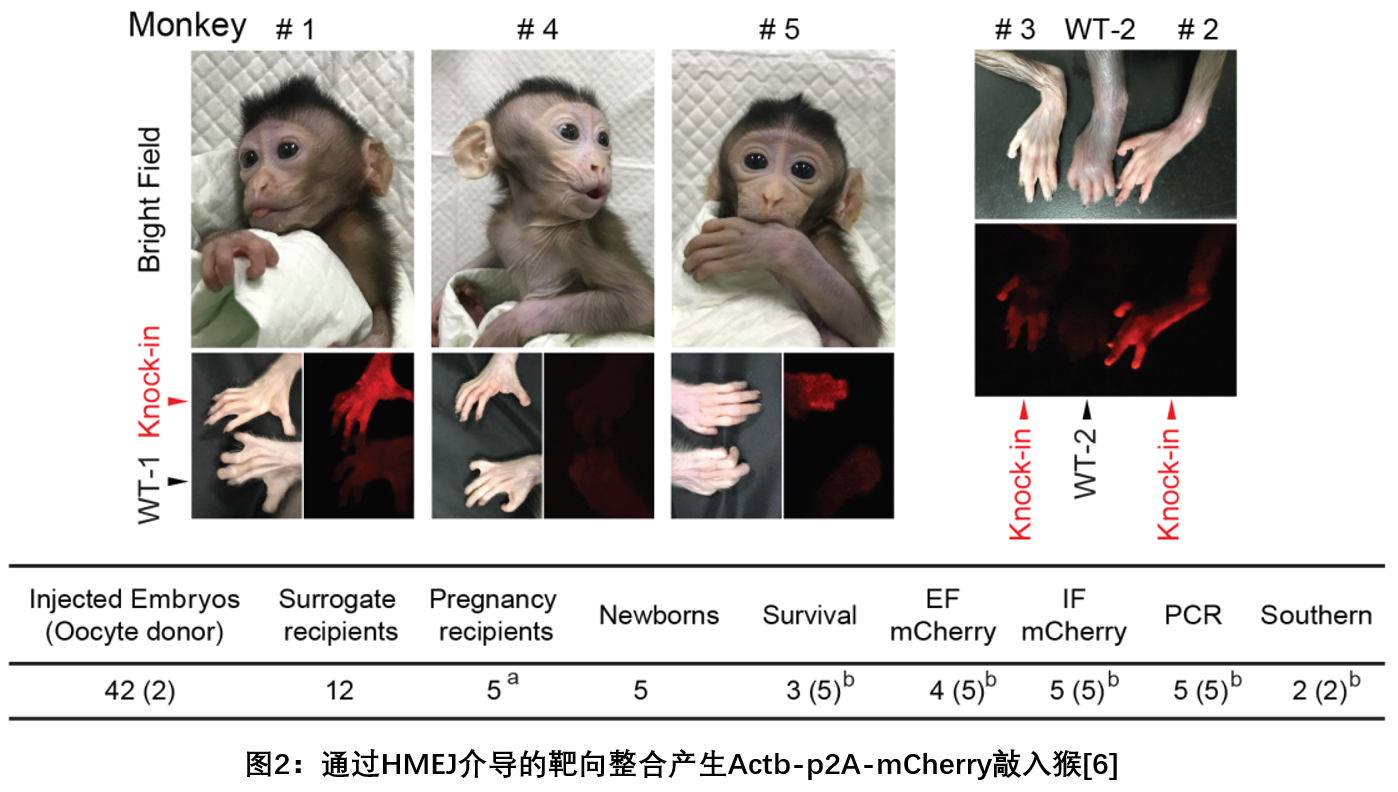
l On January 12, 2018, Cell Research magazine published two research work from Chinese scientists, reporting the birth of the world's first gene knock-in cynomolgus monkey. The Yang Hui research team from the Institute of Neuroscience of the Chinese Academy of Sciences used the previously developed homologous end-mediated integration (HMEJ) method to successfully obtain the ACTb-T2A-mcherry gene fixed-point knocking by optimizing the injection conditions by embryonic pronuclear injection. Eat crab monkeys (see Figure 2) and confirm that these monkeys have the potential for reproductive inheritance. This study provides a new solution to the construction of gene knock-in monkeys by improving integration efficiency [6].
Another study was carried out by a team of Ji Weizhi academicians who succeeded in constructing gene knockout cynomolgus monkeys using CRISPR technology in the early years. They further extended to cynomolgus monkeys based on previous techniques for precise gene editing in rats using CRISPR-Cas9. The cynomolgus monkeys with GFP integrated at the last codon of Oct4 were successfully obtained, and a few mismatches and off-targets were detected. This study demonstrates the feasibility of successful primate editing of primates and provides a versatile tool for primate reprogramming studies [7].
l On August 23, 2018, the research team of the Institute of Stem Cell and Regeneration Innovation of the Chinese Academy of Sciences published a report in Nature, which first realized the systematic knockout of SIRT6 in non-human primates, and obtained the world's first specific longevity gene knockout. The cynomolgus monkey model (see Figure 3) identifies the biological function of SIRT6 in non-primate animals. The results of the study indicate that SIRT6 is involved in the regulation of the development of non-human primates, and the lack of SIRT6 leads to delayed development of the fetus in the uterus. The establishment of this model may open up new avenues for simulating and studying the pathogenesis of human perinatal lethal syndrome [8]. 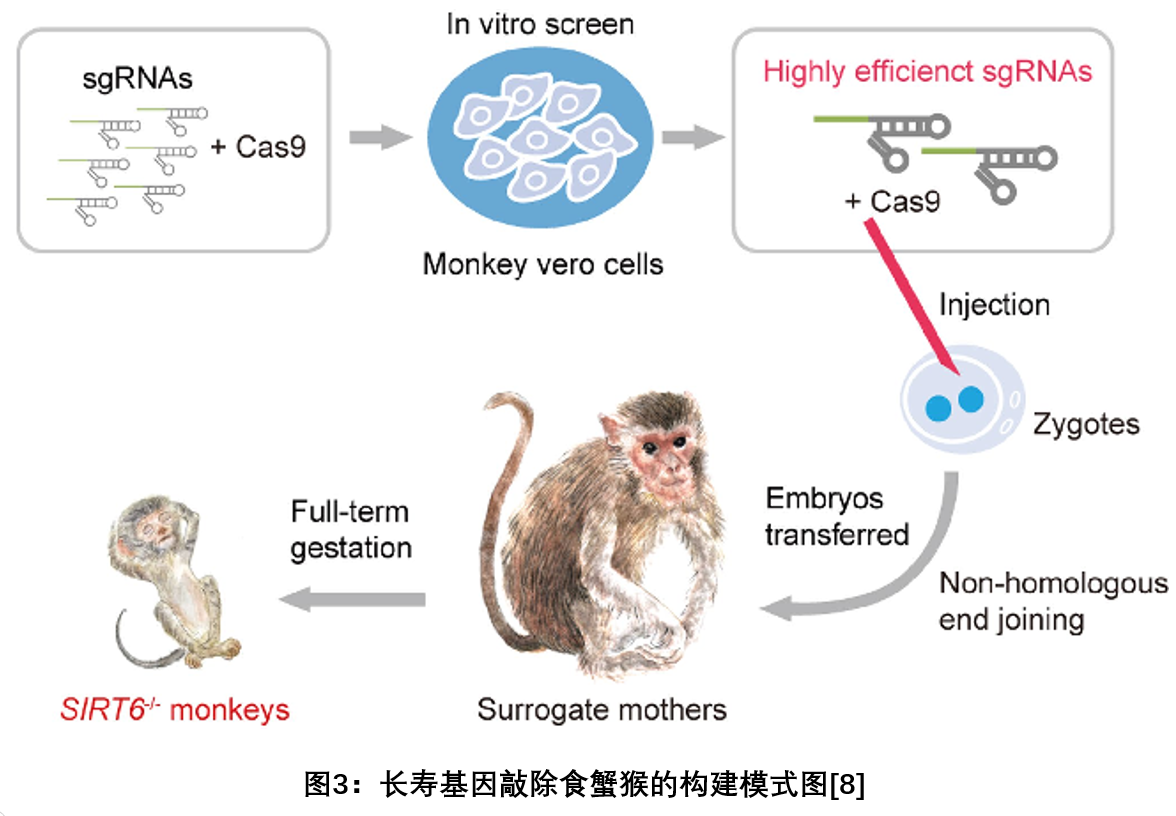
Second, the optimization of single-base gene editing technology <br> Single-base gene editing technology refers to the gene editing technology that can cause a single base change in the genome. The basic principle is the formation of cytosine deaminase (APOBEC) or adenosine deaminase fused to existing Cas9n (D10A), relying on the principle of CRISPR to make the target away from the PAM end of the 4-7 single base modified gene Editing technology. The single base editing system is the most safe and effective gene editing tool in terms of its ability to perform base substitution without cutting the genome, but its imperfect problems such as editing range, editing window, editing efficiency and specificity still need further Land transformation and optimization.
1. xCas 9: Cas9 Rapid Evolution System Expands Targeting Range of Base Editing Tools
In February 2018, David Liu's lab published a feature in Nature, using phage-assisted continuous evolutionary systems to evolve Cas9 to obtain a variant of SpCas9 (xCas9) that recognizes multiple PAMs, which can recognize NG, GAA, and GAT. Multiple PAM sequences within can target approximately one-fourth of the target in the human genome. xCas9 supports a variety of applications in human cells, including targeted transcriptional activation, nuclease-mediated gene knockout, and cytidine and adenine base editing. xCas9 has higher DNA specificity than spCas9, and when targeting non-NGG PAMs to genomic loci, xCas9 also exhibits very low non-targeting activity. These findings expand the DNA targeting range of the CRISPR system, and the fusion of ABEs also makes the selection of ABEs PAM more flexible [9].
2. dCpf1-B E s: a base editor that recognizes T-rich P AM
In May 2018, Chen Jia Laboratory of Shanghai University of Science and Technology published a report on Nature Biotechnology, which developed a CRISPR-Cpf1-based dCpf1-BEs by fusing rat cytosine deaminase APOBEC1 into catalytically inactive Cpf1. Overcoming the limitations of current BEs that can only target G/C-rich PAM sequences, enabling them to recognize T-rich PAM sequences and catalyze C-T conversion in human cells, while producing lower editing by-products, Has a higher editing accuracy. These findings provide new ideas and expand the application scope for the comprehensive and in-depth application of the base editing system [10]. 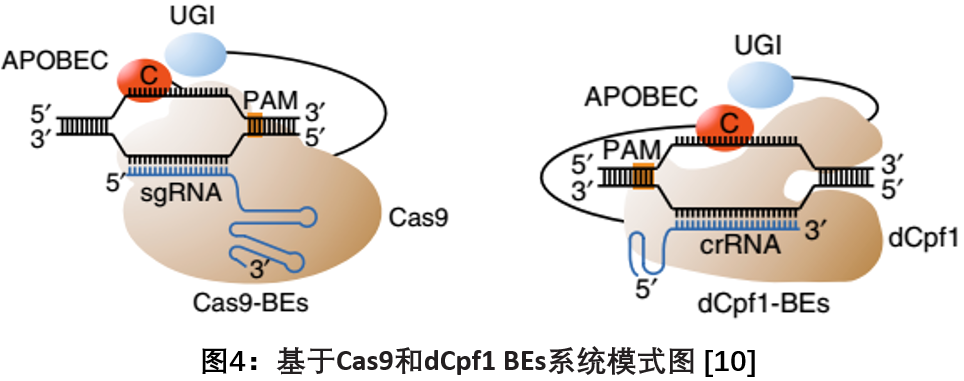
3. BE4max and ABEmax: A technical bottleneck that optimizes the base editor to address expression levels
In May 2018, David Liu Lab continued to publish in Nature Biotechnology, arguing that expression levels are a technical bottleneck in the base editor, through the introduction of nuclear localization signals (NLS) and codon optimization, and the original reconstruction of deaminase components. Optimizing the cytidine (BE4) and adenine (ABE7.10) base editors, the BE4max and ABEmax with higher editing efficiency were obtained, which were increased by 1.9 times and 1.3-7.9 times, respectively. The optimized base editor calibrates pathogenic SNPs and greatly improves editing efficiency in a variety of mammalian cells [11].
4. BE-PLUS: New base editing tool with wider editing window and higher fidelity
In July 2018, the Chen Jieping team of the Army Military Medical University and the Huang Xunxu group of the Shanghai University of Science and Technology collaborated on Cell Research to develop a new editing technique, BE-PLUS, to fuse 10 copies of the GCN4 peptide to nCas9 ( In D10A), scFv - APOBEC - UGI - GB1 was recruited to the target site, thereby editing the C to U (T) to a greater extent, greatly expanding the editing window of the base editor. The new system achieves base editing by expanding the window, thereby expanding the targeting range of the genome and improving fidelity [12]. 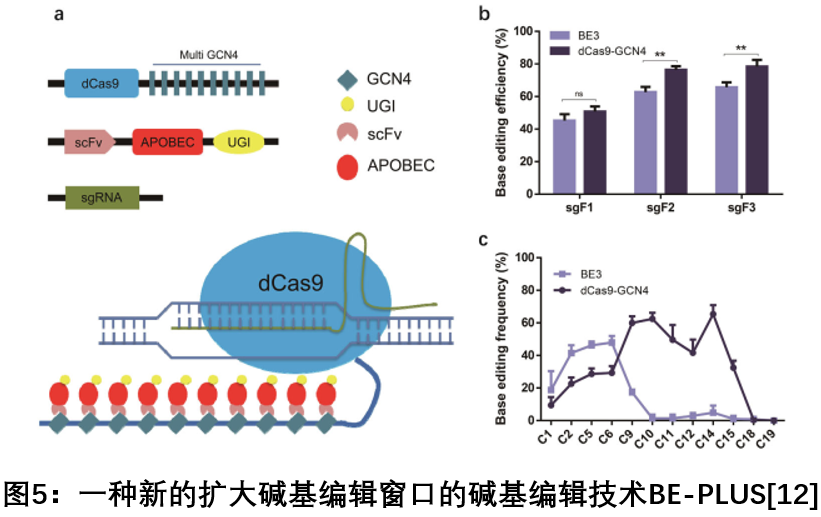
5. Optimize base editor for efficient editing in cells, organs and mice
In July 2018, Nature Biotechnology published a new study to redesign the sequences of BE3, BE4Gam and xBE3 by codon optimization and the addition of additional nuclear localization sequences. Optimized screening of constitutive and inducible base editing systems greatly increases the efficiency of C to T mutations, and the redesigned base editor can also target modifications in a variety of mouse and human cell lines as well as in intestinal organs. The study also successfully mediated effective in vivo cell editing in adult mouse liver. These optimizations help single-base editing tools to be more efficient and flexible [13].
6. eA3A-BE3: Engineered human source APOBEC3A reduces peripheral mutations at the target site
In July 2018, J. Keith Joung, a team at Harvard University and Massachusetts General Hospital, published a report on Nature biotechnology to reduce CRISPR-Cas9-based cytidine deaminase by using an engineered human APOBEC3A (eA3A) domain. A strategy for peripheral mutations that deamination of cytidine in a particular motif based on a priority of TCR > TCY > VCN, greatly reducing the presence of more than one ubiquitous five base pair editing window At C, existing base editors can create unnecessary C to T conversion possibilities. eA3A-BE3 shows higher precision (more than 40 times) and lower off-target efficiency than BE3, and maintains similar editing activity. Using this system, they also efficiently calibrated mutations in the human beta-thalassemia promoter, providing a more accurate editing tool for the clinical application of base editing [14]. 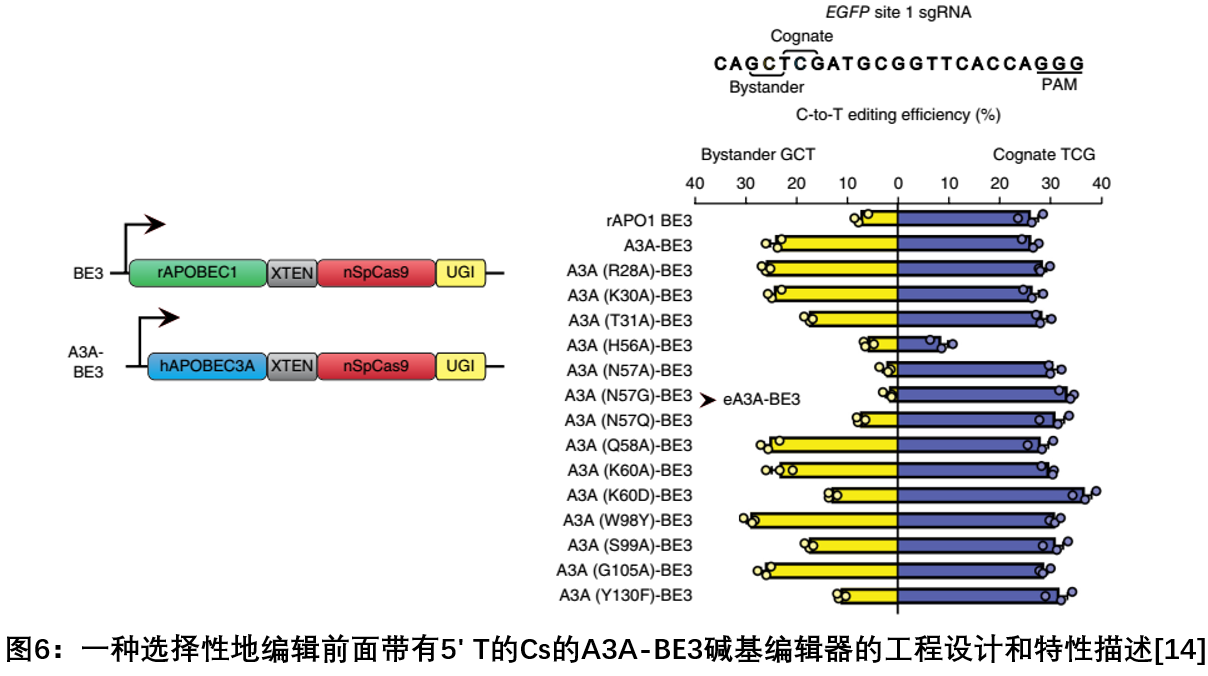
7. hA3A-BE3: : Human APOBEC3A enables efficient editing of highly methylated regions C to T
In August 2018, researchers from the Shanghai University of Science and Technology and the Chinese Academy of Sciences published a report in Nature Biotechnology. By screening multiple APOBEC and AID deaminase, it was found that the human APOBEC3A binding to BEs has a narrower editing window at the methylation level and The region with high GpC dinucleotide content efficiently mediates C to T base editing, solving the problem that the current APOBEC1-based BEs system is relatively inefficient in editing highly methylated cytosines [15]. 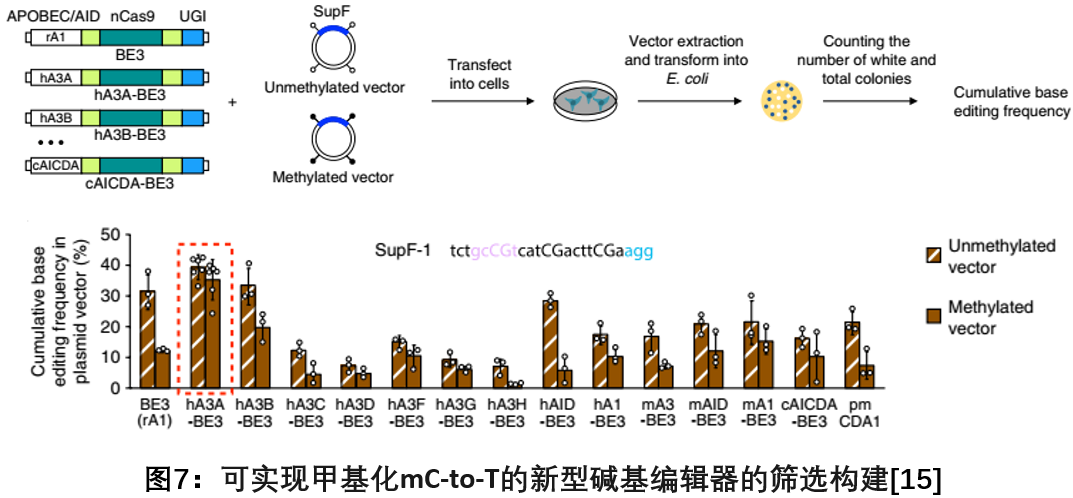
8. In the same period, two reports used the A BE s system to construct the mouse disease model for the first time .
In September 2018, the Protein & Cell journal published two highly efficient base editing in mice and rats using the Optimized Adenine Base Editor (ABE). The paper not only highlights the fidelity and efficiency of ABEs in the induction of A-T to G-C, but also demonstrates their potential ease of use in generating disease models and correcting disease mutations in animal and human embryos. Feature. details as follows:
l Professor Liu Mingyao, Dean of the School of Life Sciences of East China Normal University, and Professor Liu Mingyao, the chief scientist of Bangyao Biotechnology, and the team member Professor Li Dali, created an effective ABE system to effectively create three mouse strains and successfully constructed hereditary muscle degeneration. Mouse model of (DMD). In addition, a rat model of type II hereditary glycogen storage disease (GSD) was constructed. This is the first report to achieve efficient point mutations in mice through a base editing system. This research not only has great potential in the construction of disease models, but more importantly, it has important application value in the treatment of hereditary diseases caused by point mutations [16].
l Huang Jun of Sun Yat-sen University and the Songyangzhou team also tested the effectiveness of the ABEs system in the construction of disease mouse models. Mouse albino and DMD models were successfully constructed by targeting ABE to target mRNA splice sites to induce gene dysfunction. These results indicate that ABEs can efficiently and accurately convert base A in mouse embryos to G, demonstrating that this is a high-fidelity tool for generating point-mutated mouse models [17].
9. A 3A-PBE : Efficient C to T editing in plants using the fusion of nCas9 and human APOBEC3A
In October 2018, Gao Caixia, a research group of the Institute of Genetics and Developmental Biology, Chinese Academy of Sciences, published a report on Nature Biotechnology, showing a base editing system (A3A-PBE) consisting of Cas9 nickase and human APOBEC3A (A3A-PBE). Efficient C to T base editing in wheat, rice and potato. At the same time, the uracil glycosylase inhibitor (UGI) and MU were fused to A3A-PBE, and the A3A-Gam system was created, which greatly improved the efficiency of base editing. The new system not only expands the 5 nt sequence editing window based on rat APOBEC1 to 17 nt, but also effectively edits areas with high GC content, which has important application value for new variety cultivation and crop genetic improvement [18]. 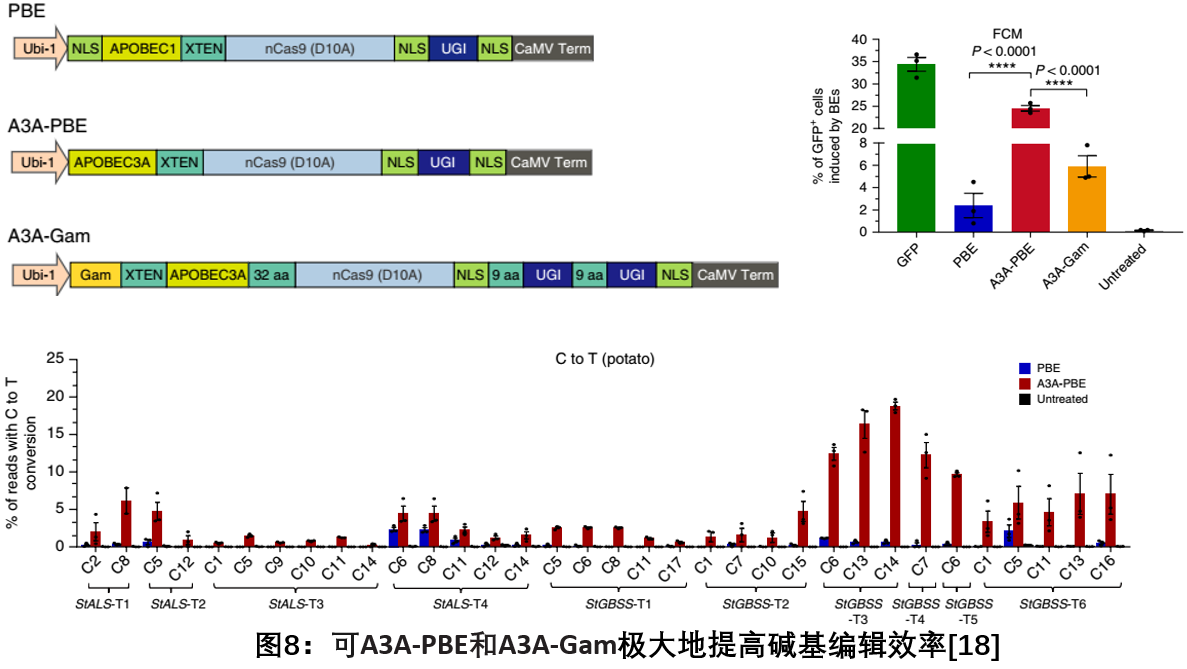
Summary <br> In summary, we briefly introduced the breakthrough of the 2018 CRISPR system in large animal model construction and single base editing technology. Looking ahead, we can foresee the rapid development of the basic research and clinical research that CRISPR-Cas9 technology will bring. Although not fully developed at this stage, this technology has revolutionized biological and medical research, including genome editing, gene screening and cell tracking, disease treatment and more. For more important applications, we will continue to introduce them in the next issue!
references:
[1] http:// .
[2] Yan, Sen, et al. "A Huntingtin Knockin Pig Model Recapitulates Features of Selective Neurodegeneration in Huntington ' s Disease." Cell (2018): S009286741830285X.
[3] Liu, Zhen, et al. "Cloning of Macaque Monkeys by Somatic Cell Nuclear Transfer." Cell (2018): S0092867418300576.
[4] Peiyuan Qiu , et al. " BMAL1 knockout macaque monkeys display reduced sleep and psychiatric disorders. " National Science Review (2019): https://doi.org/10.1093/nsr/nwz002 .
[5] Zhen Liu , et al. " Cloning of a gene-edited macaque monkey by somatic cell nuclear transfer. " National Science Review (2019): https://doi.org/10.1093/nsr/nwz003 .
[6] Yao, Xuan, et al. "Generation of knock-in cynomolgus monkey via CRISPR/Cas9 editing." Cell Research (2018).
[7] Cui, Yiqiang, et al. "Generation of a precise Oct4-hrGFP knockin cynomolgus monkey model via CRISPR/Cas9-assisted homologous recombination." Cell Research 28.3 (2018).
[8] Weiqi, Zhang, et al. "SIRT6 deficiency results in developmental retardation in cynomolgus monkeys." Nature (2018).
[9] Hu, Johnny H., et al. "Evolved Cas9 variants with broad PAM compatibility and high DNA specificity." Nature.
[10] Li, X. et al. Base editing with a Cpf1-cytidine deaminase fusion. Nat Biotechnol190 36, 324-327 (2018)
[11] Koblan, Luke W, et al. "Improving cytidine and adenine base editors by expression optimization and ancestral reconstruction." Nature Biotechnology (2018).
[12] Wen, Jiang, et al. "BE-PLUS: a new base editing tool with broadened editing window and enhanced fidelity." Cell Research (2018).
[13] Paz, Zafra Maria, et al. "Optimized base editors enable efficient editing in cells, organoids and mice." Nature Biotechnology (2018).
[14] Gehrke, Jason M, et al. "An APOBEC3A-Cas9 base editor with minimized bystander and off-target activities." Nature Biotechnology (2018).
[15] Xiao, Wang, et al. "Efficient base editing in methylated regions with a human APOBEC3A-Cas9 fusion." Nature Biotechnology (2018).
[16] Yang, Lei, et al. "Increasing targeting scope of adenosine base editors in mouse and rat embryos through fusion of TadA deaminase with Cas9 variants." Protein & Cell 9.9 (2018).
[17] Liang, Puping, et al. "Effective and precise adenine base editing in mouse zygotes." Protein & Cell 9.9 (2018).
[18] Zong Y , et al. Efficient C-to-T base editing in plants using a fusion of nCas9 and human APOBEC3A. Nature Biotechnology (2018).
Foreword : In recent years, CRISPR gene editing technology is sweeping the entire field of biomedical research. In the previous issue, we have sorted out the 2018 related events from the aspects of CRISPR system development and mechanism research. With the continuous optimization of basic technologies, the application of CRISPR technology is also more extensive, such as animal modeling, drug screening, single-base editing technology, cell lineage tracing, basic disease research, disease diagnosis, in vivo editing and genetic disease correction. It is worth mentioning that the world's first CRISPR-based gene editing clinical trial was implemented in 2018 [1]. In December of the same year, the FDA approved an application for clinical trial of Editas using CRISPR technology to treat congenital black shank disease. It is expected to become the first therapy in the world to use CRISPR technology in the human body. In this issue, we will take a look at the major breakthroughs in the application of CRISPR technology in the past year from the aspects of animal modeling and single-base technology.